Biomodels
The BioModels database is a repository of peer-reviewed, published, computational models suitable for simulating biological processes.
These mathematical models are primarily from the field of systems biology, but more generally of biological interest. This resource allows biologists to store, search and retrieve published mathematical models. In addition, models in the database can be used to generate sub-models, can be simulated online, and can be converted between different representational formats. This resource also features programmatic access via Web Services.
The BioModels database is developed and maintained by the BioModels.net initiative.
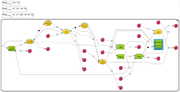
BioUML provides possibility to visualize and browse the diagrams and models from the Biomodels database. The database can be found in the repository pane with read access only. The Diagrams subdirectory currently contains 421 curated and 433 non-curated diagrams. To find a certain model you can either scroll down and (in the web edition) subsequently expand the subdirectory as necessary or just run a text search through the database.
When a diagram is opened in the document pane, the Info tab in the information box shows some details about the model, including database links for the individual components. In the viewparts area, under the tab My description a detailed description of the model is given; it shows the original reference, its abstract and the PubMed link; further information about how the model was constructed is added.
Under the Simulation tab you will find the default settings for the simulation, which can be changed before launching the simulation (![]() ). The simulation results will be graphically displayed in a new window, which can be saved as an image (in a number of formats) using the Export button (
). The simulation results will be graphically displayed in a new window, which can be saved as an image (in a number of formats) using the Export button (![]() ) in the general control panel.
) in the general control panel.