Diagram type
From BioUML platform
The diagram type concept was introduced to take into account different diagram types and problem domain specificity. The diagram type defines:
- types of biological components and their interactions that can be shown on the diagram;
- diagram view builder – it generates view (image) for each graph element taking into account the problem domain peculiarities. For example, biological pathway diagram view builder displays proteins as circles, genes as rectangles and substances as squares;
- semantic controller - provides semantic integrity of the diagram during its editing. It takes into account problem domain constraints, for example if some substance is removed on biological pathway diagram, all related reactions must be removed too.
A diagram type can be defined (created) by two ways:
- programmatically - as Java class implementing special interface. There are 5 predefined diagram types that allow to describe complex biological systems on cellular level with different level of detail and formality;
- declaratively - as XML document. BioUML workbench provides Graphic Notation Editor that allows advanced users to create and edit diagram types. By this way diagram types for SBGN (Le Novère et al, 2009) and KEGG/Pathways database were created.
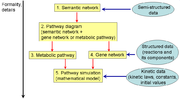
BioUML provides a number of diagram types that allow a researcher to describe biological pathways such as metabolic pathways, signal transduction pathways and gene networks. The main three of them are:
- Semantic network is used to describe semantic relationships between system components, system states, and related problem domain concepts. This diagram type is also convenient as overview.
- Pathway is used for formalized description of biological pathway structure (metabolic pathway or gene network).
- Pathway simulation is an extension of the pathway type, where variables are associated with graph nodes and differential equations - with graph edges. This allows BioUML to automatically generate a mathematical model of the system and simulate its dynamics.
- Mathematical model is mathematically equivalent to pathway simulation. Which means that any ODE/algebraic mathematical model created in Pathway simulation may be also created using this diagram type. It does not contain specifical to biochemical models reactions and entities and can not be simulated with stochastic engine.
- Composite model - Hierarchical diagram containing other diagrams as parts (subdiagrams). Subdiagrams may be one of these types: SBML-SBGN, Pathway simulation or Mathematical model. Before numerical simulation is performed, such diagram is automatically flattened into corresponding ODE/algebraic diagram.
- Agent-based model - hierrachical diagram supporting arbitrary diagrams (not only ODE/algebraic). Simulation is performed for each subdiagram separately with its own simulation engine, scheduler coordinates signal exchange bewteen them.
- SBML in SBGN notation - combines SBGN visual representation with SBML mathematics. Each SBML object has corresponding visual glyph. For equations and events we use non-SBGN glyphs.
- SBML comp in SBGN notation - hierarchical diagram containing other SBML-SBGN models as elements (subdiagrams). Is is automatically flattened into SBML-SBGN diagram before numerical calculations.
- Arterial tree - describes structure and properties (areas and elasticities) of vessel bed. Numerical simulations provides information about blood flow (pressures, flow rates) across given vessel bed.
- Population-based model - contains nonlinear mixed effects (nlme) mode. It contains reference to another diagram which is used as structural model (usually ODE).
- BioNetGen - visual representation for the model of BioNetGen rules.
-