Difference between revisions of "Tree-table (element type)"
Tagir Valeev (Talk | contribs) (Created page with "{{Element type|tree-table|ru.biosoft.treetable.TreeTableElement}} === Description === This element represents a mixture of table and the tree: each tree branch or leaf can be ...") |
Tagir Valeev (Talk | contribs) |
||
| Line 8: | Line 8: | ||
Internally this element has the following properties: | Internally this element has the following properties: | ||
* '''TreePath''': path to some collection from the [[repository]] which represents a tree root; | * '''TreePath''': path to some collection from the [[repository]] which represents a tree root; | ||
| − | * '''TableScript''': JavaScript code which is used to generate a {{Type link|table}}. If table from the repository should be used, then you may use the following code: | + | * '''TableScript''': [[Using JavaScript in BioUML|JavaScript]] code which is used to generate a {{Type link|table}}. If table from the repository should be used, then you may use the following code: |
[[data (host object)|data]].get("<path_to_table>"); | [[data (host object)|data]].get("<path_to_table>"); | ||
Element from the tree is annotated using the table row with the same name. | Element from the tree is annotated using the table row with the same name. | ||
Latest revision as of 13:30, 2 July 2013
[edit] Description
This element represents a mixture of table and the tree: each tree branch or leaf can be annotated using the data from table columns. This element can be opened as a tree-table document (default) or a table document (in this case only the corresponding table will be displayed). To open as table use ![]() Open table context action in the repository pane.
Open table context action in the repository pane.
Currently such item cannot be imported, exported or produced by any analysis. You may find tree-tables in GTRD database in views collection.
[edit] Implementation
Internally this element has the following properties:
- TreePath: path to some collection from the repository which represents a tree root;
- TableScript: JavaScript code which is used to generate a
 table. If table from the repository should be used, then you may use the following code:
table. If table from the repository should be used, then you may use the following code:
data.get("<path_to_table>");
Element from the tree is annotated using the table row with the same name.
- HideBranchesAbsentInTable: if true then branches absent in the table will not be displayed (default: false).
[edit] File format
There's a transformer class FileViewTransformer which allows to load tree-table from the file. An example of such file is displayed here:
[TreeTable]
TreePath=databases/GTRD/Dictionaries/classification
HideAbsentBranches = false
[Script]
var query = "select c.name TF,c.title Title,"+
"if(e.id is null,(select concat('[\"',count(1),'\"]') from chip_experiments where tfClassId like concat(c.name,'.%')),"+
"concat('[\"',group_concat(e.id separator '\",\"'),'\"]')) Experiments, "+
"concat('[\"',group_concat(h.input separator '\",\"'),'\"]') Peaks "+
"from classification c left join chip_experiments e on e.tfClassId=c.name "+
"left join hub h on h.output=e.id "+
"where h.input_type is null or (h.input_type='PeaksGTRDType' and h.output_type='ExperimentGTRDType') group by c.name";
var t=table.query("databases/GTRD", query, "$1");
t.getColumnModel().getColumns()[1].setValue("referenceType", "ChIP-seq peaks: GTRD");
t.getColumnModel().getColumns()[1].setType(Packages.ru.biosoft.table.TableColumn.DataType.Set);
t.getColumnModel().getColumns()[2].setValue("referenceType", "ChIP experiment: GTRD");
t.getColumnModel().getColumns()[2].setType(Packages.ru.biosoft.table.TableColumn.DataType.Set);
t;
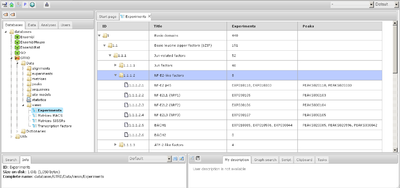
This tree-table looks in BioUML web edition like here: