Difference between revisions of "Parameter identifiability example"
(→Interpretation of results) |
|||
| (35 intermediate revisions by one user not shown) | |||
| Line 1: | Line 1: | ||
<font size = 3> | <font size = 3> | ||
| − | Identifiability analysis infers how well the model parameters are approximated by the amount and quality of experimental data [1,2]. | + | Identifiability analysis infers how well the model parameters are approximated by the amount and quality of experimental data [1,2]. For this, we consider the optimization problem of minimizing the distance function, defined as a normalized sum of squared differences between the experimental and simulated values of the observed variables. Identifiability analysis seeks a solution to this problem at a fixed value of each of the fitting parameters in turn, and examines the behavior of the objective function as the value increases (steps to the right) and decreases (steps to the left). There are three options depending on whether the objective function value exceeds the allowable threshold with the changing parameter value: |
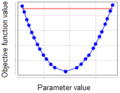
| + | *exceeds in both directions, then the parameter is ''identifiable'' (Figure 1); | ||
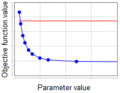
| + | *exceeds only in one direction, then the parameter is ''partially non-identifiable'' (Figure 2); | ||
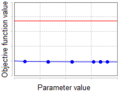
| + | *does not exceed, then the parameter is ''non-identifiable'' (Figure 3). | ||
| + | |||
| + | <font size=2> | ||
| + | <gallery> | ||
| + | File:parameter_identifiability_figure_1.png|'''Figure 1''': Profile of the identifiable parameter. | ||
| + | File:parameter_identifiability_figure_2.png|'''Figure 2''': Profile of the partially non-identifiable parameter. | ||
| + | File:parameter_identifiability_figure_3.png|'''Figure 3''': Profile of the non-identifiable parameter. | ||
| + | </gallery> | ||
| + | </font> | ||
==Reproducing a test case in BioUML== | ==Reproducing a test case in BioUML== | ||
| − | To reproduce a test case | + | To reproduce a test case in the [[BioUML]] workbench, go to the <b>Analyses</b> tab in the navigation pane and follow to:<br> ''analyses'' > ''Methods'' > ''Differential algebraic equations''. |
Identifiability analysis can be run in two ways: | Identifiability analysis can be run in two ways: | ||
| Line 9: | Line 20: | ||
*for auto-generation of an optimization document using the given settings, double click on '''Parameter identifiability (table)'''. | *for auto-generation of an optimization document using the given settings, double click on '''Parameter identifiability (table)'''. | ||
| − | < | + | ==Parameter identifiability (optimization)== |
| + | |||
| + | Definition of the method parameters can be found [[Parameter identifiability (optimization) (analysis)|here]]. | ||
| + | |||
| + | For our example, we used a test case optimization created for the MAP kinase cascade model of Kholodenko [3]. A brief description of this test case is done in the chapter [[Optimization examples]]. In the current identifiability analysis, we used the following settings: | ||
| + | |||
| + | <ul> | ||
| + | <li>Path to an optimization document in the BioUML repository:<br> | ||
| + | '''Optimization''' = ''data/Examples/Optimization/Data/Documents/test_case_2'' | ||
| + | |||
| + | <li>Path to an optimization results:<br> | ||
| + | '''Parameter values''' = ''data/Examples/Optimization/Data/Simulations/optimization_results_2/optimizationInfo'' | ||
| + | |||
| + | <li>The maximum number of steps performed by the analysis for each fitting parameter in one direction:<br> | ||
| + | '''Maximum identifiability steps''' = 50 | ||
| + | |||
| + | <li>As the maximal deviation from the initial objective function value (allowable threshold), we considered ten percent of the smallest objective function value (found by the optimization and corresponding to the results in the ''optimizationInfo'' table):<br> | ||
| + | '''Maximal deviation''' = 4.2 | ||
| + | |||
| + | <li>A possible path to save results of the analysis:<br> | ||
| + | '''Output path''' = ''data/Collaboration/Demo/Data/Temp/Identifiability results (for test_case_2)'' | ||
| + | </ul> | ||
<br>[[File:parameter_identifiability_example_01.png]]<br><br> | <br>[[File:parameter_identifiability_example_01.png]]<br><br> | ||
| − | < | + | After you click the <b>Run</b> button and the calculations are finished, you will see a result table that includes information on all fitting parameters of the optimization: parameter names (the column ''"Name"''), start values (''"Value"''), values improved by the analysis (''"Estimated value"'', if these values are equal to the start values, the analysis could not improve the solution of the optimization problem, i.e. started with the best solution), objective function values for the estimated values (''"Objective function value"''), and links to images, showing the identifiability profile of parameters (''"Plot path"''): |
| + | |||
| + | <br>[[File:parameter_identifiability_example_05.png]]<br><br> | ||
| + | |||
| + | Clicking on any plot path link will open a new tab with the identifiability profile of the corresponding parameter: | ||
| + | |||
| + | <br>[[File:parameter_identifiability_example_06.png]]<br><br> | ||
| + | |||
| + | Ready analysis results can be found in the folder: | ||
| + | |||
| + | ''data/Examples/DAE models/Data/Parameter Identifiability'' | ||
| + | |||
| + | ==Parameter identifiability (table)== | ||
| + | |||
| + | Definition of the method parameters can be found [[Parameter identifiability (table) (analysis)|here]]. | ||
| + | |||
| + | To get results described above, firstly set the following options: | ||
| + | |||
| + | <ul> | ||
| + | <li>Path to a diagram utilized in the ''test_case_2'' optimization:<br> | ||
| + | '''Diagram''' = ''data/Examples/Optimization/Data/Diagrams/diagram_2'' | ||
| + | |||
| + | <li>Path to a table with the same fitting parameters as in the ''test_case_2'' optimization:<br> | ||
| + | '''Parameters table''' = ''data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Parameters table''<br> | ||
| + | As the start values (the column ''"Val"''), this table contains the best solution of the optimization problem: | ||
| + | |||
| + | <br><br>[[File:parameter_identifiability_parameters_table.png]]<br><br> | ||
| + | |||
| + | <li>Path to a table with experimental data:<br> | ||
| + | '''Observables table''' = ''data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Observables table''<br> | ||
| + | Column names in the table must have the same identifiers as the corresponding entities in the model: | ||
| + | |||
| + | <br>[[File:parameter_identifiability_observables_table.png]]<br><br> | ||
| + | |||
| + | <li>Name of the time column in the ''Observbles table'':<br> | ||
| + | '''Time column''' = ''time'' | ||
| + | |||
| + | <li>The maximum number of steps performed by the analysis for each fitting parameter in one direction:<br> | ||
| + | '''Maximum identifiability steps''' = 50 | ||
| + | |||
| + | <li>The maximal deviation from the initial objective function value is the same as in the example above:<br> | ||
| + | '''Maximal deviation''' = 4.2 | ||
| + | |||
| + | <li>Optimization method parameters (the same as in the ''test_case_2'' optimization):<br> | ||
| + | '''Optimization method name''' = ''Particle swarm optimization''<br> | ||
| + | '''Number of itirations''' = 200<br> | ||
| + | '''Number of particles''' = 50 | ||
| + | |||
| + | <li>A possible path to save results of the analysis:<br> | ||
| + | '''Output path''' = ''data/Collaboration/Demo/Data/Temp/Identifiability results (for diagram_2)'' | ||
| + | </ul> | ||
| + | |||
| + | After setting these options, your workspace will look like this: | ||
| + | |||
| + | <br>[[File:parameter_identifiability_example_02.png]]<br><br> | ||
| + | |||
| + | Then click on the phrase "<u>Show expert options >></u>": | ||
| + | |||
| + | <br>[[File:parameter_identifiability_example_03.png]]<br><br> | ||
| + | |||
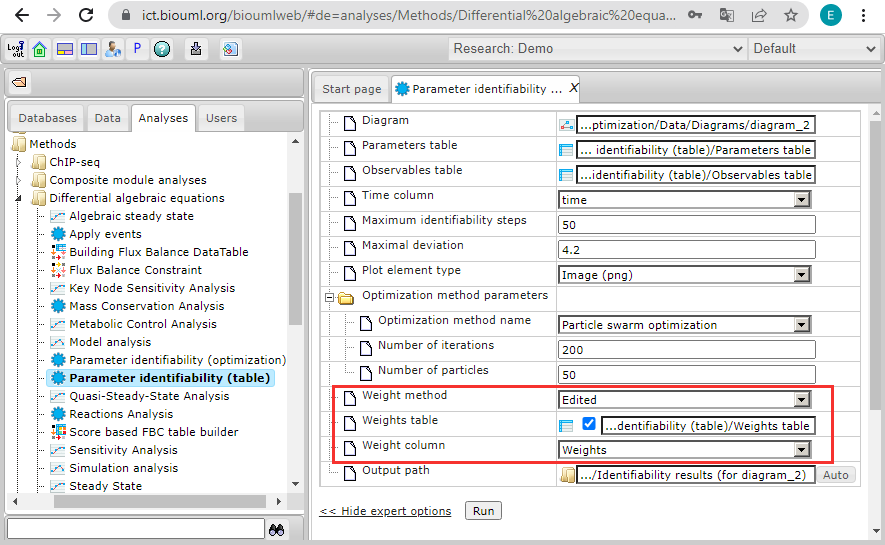
| + | And additionally set the options for complete coincidence with the optimization problem ''test_case_2'': | ||
| + | |||
| + | <ul> | ||
| + | <li>The method to estimate the objective function weights:<br> | ||
| + | '''Weight method''' = ''Edited'' | ||
| + | |||
| + | <li>Path to a table defining the required weights:<br> | ||
| + | '''Weights table''' = ''data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Weights table''<br> | ||
| + | |||
| + | <br>[[File:parameter_identifiability_weights_table.png]]<br><br> | ||
| + | |||
| + | <li>Name of the column with weights in the ''Weights table'':<br> | ||
| + | '''Weight column''' = ''Weights'' | ||
| + | </ul> | ||
| + | |||
| + | <br>[[File:parameter_identifiability_example_04.png]]<br><br> | ||
| + | |||
| + | To start the analysis, click on the <b>Run</b> button. The results table is similar to the one obtained from the analysis using the optimization document. | ||
| + | |||
| + | ==Interpretation of results== | ||
| + | In our example, we received the following identifiability profiles: | ||
| + | |||
| + | <br>[[File:parameter_identifiability_results.png]]<br><br> | ||
| + | |||
| + | Identifiability profiles for parameters ''J0_V1'', ''J1_V2'', ''J4_V5'', ''J8_V9'' cross the threshold of a given deviation from the initial objective function value in both directions. Therefore, the analysis defined these variables as identifiable. The identifiability profiles for parameters ''J5_V6'' and ''J9_V10'' cross the threshold in only one direction, so the analysis determined these parameters as partially non-identifiable. However, it should be noted that the resulting ''J5_V6'' value in the optimization problem is almost equal to the upper bound of the given search space. Thus, the identifiability analysis tried to take very small steps to the right in the parameter value, but this was not enough to significantly change the objective function value: | ||
| + | |||
| + | <code> | ||
| + | INFO - (Step 1 of 50) J5_V6 = 1.4249999525529151 (step size = 9.683078561195658E-10), objective function = 42.25511865896442<br> | ||
| + | INFO - (Step 2 of 50) J5_V6 = 1.4249999614126978 (step size = 8.859782795778178E-9), objective function = 42.25511100723304<br> | ||
| + | INFO - (Step 3 of 50) J5_V6 = 1.4249999710878445 (step size = 9.675146601603706E-9), objective function = 42.25509033270342<br> | ||
| + | INFO - (Step 4 of 50) J5_V6 = 1.4249999807495213 (step size = 9.66167670891121E-9), objective function = 42.25508972805858<br> | ||
| + | INFO - (Step 5 of 50) J5_V6 = 1.4249999904319726 (step size = 9.682451299316246E-9), objective function = 42.255089526361225<br> | ||
| + | INFO - (Step 6 of 50) J5_V6 = 1.425 (step size = 9.682869310218694E-9), objective function = 42.25525810172391<br> | ||
| + | INFO - Parameter bound exceeded. | ||
| + | </code> | ||
| + | |||
| + | An increase in the ''J9_V10'' value (steps to the right) gave a stable increase in the objective function value. However, the specified threshold was not exceeded because the parameter value reached the upper bound of the search space. Thus, changing the boundaries of the search space in the optimization problem can change the identifiability profile of model parameters. | ||
| + | |||
| + | It is also worth noting that the step sizes automatically determined by the algorithm, in particular, depend on how close the value of the fitting parameter is to the search boundary. As can be seen from the identifiability profiles above, for parameters ''J5_V6'', ''J8_V9'', and ''J9_V10'', steps to the right are much smaller than steps to the left. An increase in the upper search bounds for these parameters leads to finding a new solution to the optimization problem (the value of the objective function is 31.7 versus 42.2). In this case, all parameters become identifiable: | ||
| + | |||
| + | <br>[[File:parameter_identifiability_results_02.png]]<br><br> | ||
==References== | ==References== | ||
# Raue A, Kreutz C, Maiwald T, Bachmann J, Schilling M, Klingmüller U, Timmer J (2009) Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics, 25(15):1923–1929. | # Raue A, Kreutz C, Maiwald T, Bachmann J, Schilling M, Klingmüller U, Timmer J (2009) Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics, 25(15):1923–1929. | ||
# Raue A, Becker V, Klingmüller U, Timmer J (2010) Identifiability and observability analysis for experimental design in nonlinear dynamical models. Chaos, 20(4):045105. | # Raue A, Becker V, Klingmüller U, Timmer J (2010) Identifiability and observability analysis for experimental design in nonlinear dynamical models. Chaos, 20(4):045105. | ||
| + | # Kholodenko BN (2000) Negative feedback and ultrasensitivity can bring about oscillations in the mitogenactivated protein kinase cascades. European Journal of Biochemistry, 267(6):1583–1588. | ||
</font> | </font> | ||
Latest revision as of 14:09, 18 March 2022
Identifiability analysis infers how well the model parameters are approximated by the amount and quality of experimental data [1,2]. For this, we consider the optimization problem of minimizing the distance function, defined as a normalized sum of squared differences between the experimental and simulated values of the observed variables. Identifiability analysis seeks a solution to this problem at a fixed value of each of the fitting parameters in turn, and examines the behavior of the objective function as the value increases (steps to the right) and decreases (steps to the left). There are three options depending on whether the objective function value exceeds the allowable threshold with the changing parameter value:
- exceeds in both directions, then the parameter is identifiable (Figure 1);
- exceeds only in one direction, then the parameter is partially non-identifiable (Figure 2);
- does not exceed, then the parameter is non-identifiable (Figure 3).
Contents |
[edit] Reproducing a test case in BioUML
To reproduce a test case in the BioUML workbench, go to the Analyses tab in the navigation pane and follow to:
analyses > Methods > Differential algebraic equations.
Identifiability analysis can be run in two ways:
- to use a pre-created optimization document, double click on Parameter identifiability (optimization);
- for auto-generation of an optimization document using the given settings, double click on Parameter identifiability (table).
[edit] Parameter identifiability (optimization)
Definition of the method parameters can be found here.
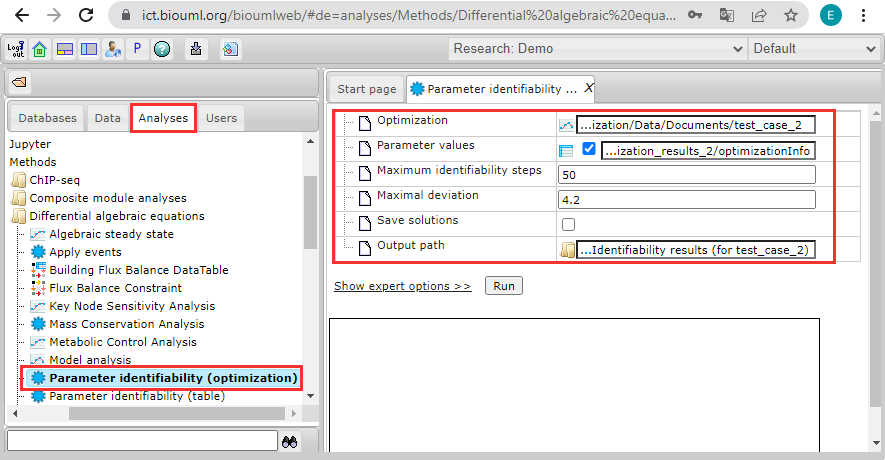
For our example, we used a test case optimization created for the MAP kinase cascade model of Kholodenko [3]. A brief description of this test case is done in the chapter Optimization examples. In the current identifiability analysis, we used the following settings:
- Path to an optimization document in the BioUML repository:
Optimization = data/Examples/Optimization/Data/Documents/test_case_2 - Path to an optimization results:
Parameter values = data/Examples/Optimization/Data/Simulations/optimization_results_2/optimizationInfo - The maximum number of steps performed by the analysis for each fitting parameter in one direction:
Maximum identifiability steps = 50 - As the maximal deviation from the initial objective function value (allowable threshold), we considered ten percent of the smallest objective function value (found by the optimization and corresponding to the results in the optimizationInfo table):
Maximal deviation = 4.2 - A possible path to save results of the analysis:
Output path = data/Collaboration/Demo/Data/Temp/Identifiability results (for test_case_2)
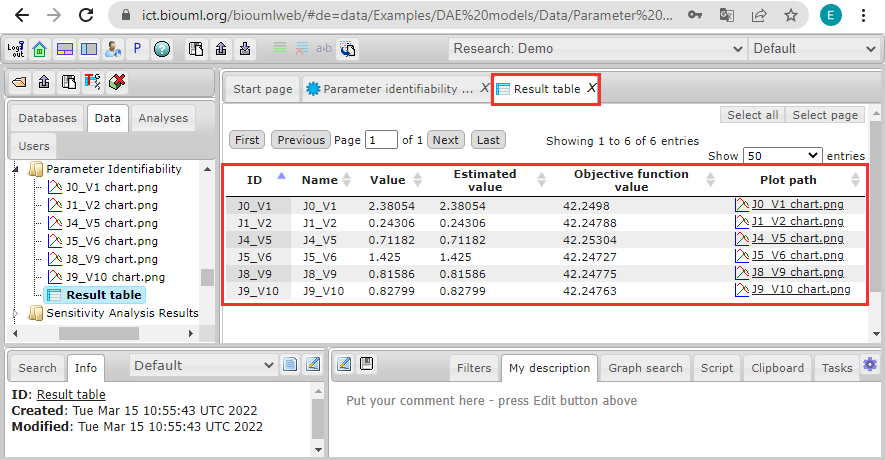
After you click the Run button and the calculations are finished, you will see a result table that includes information on all fitting parameters of the optimization: parameter names (the column "Name"), start values ("Value"), values improved by the analysis ("Estimated value", if these values are equal to the start values, the analysis could not improve the solution of the optimization problem, i.e. started with the best solution), objective function values for the estimated values ("Objective function value"), and links to images, showing the identifiability profile of parameters ("Plot path"):
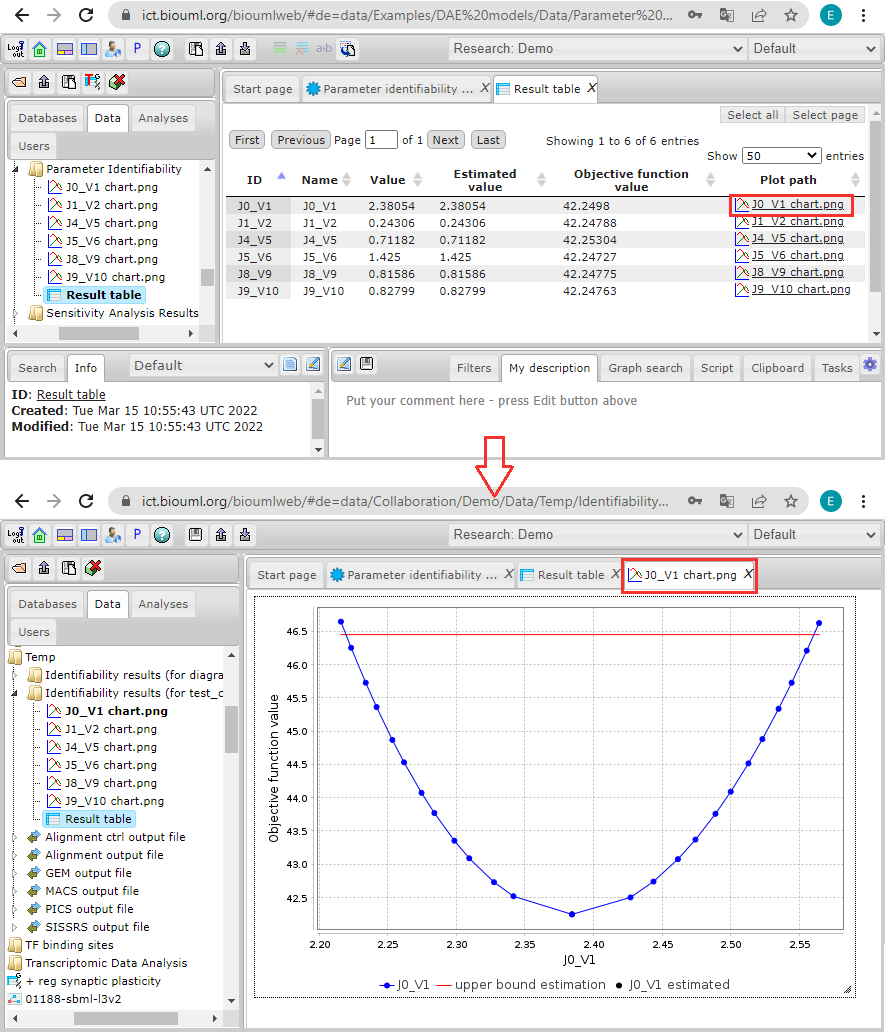
Clicking on any plot path link will open a new tab with the identifiability profile of the corresponding parameter:
Ready analysis results can be found in the folder:
data/Examples/DAE models/Data/Parameter Identifiability
[edit] Parameter identifiability (table)
Definition of the method parameters can be found here.
To get results described above, firstly set the following options:
- Path to a diagram utilized in the test_case_2 optimization:
Diagram = data/Examples/Optimization/Data/Diagrams/diagram_2 - Path to a table with the same fitting parameters as in the test_case_2 optimization:
Parameters table = data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Parameters table
As the start values (the column "Val"), this table contains the best solution of the optimization problem: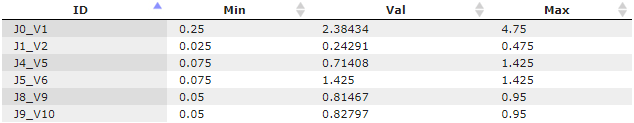
- Path to a table with experimental data:
Observables table = data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Observables table
Column names in the table must have the same identifiers as the corresponding entities in the model: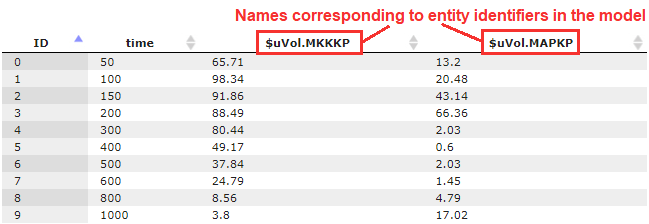
- Name of the time column in the Observbles table:
Time column = time - The maximum number of steps performed by the analysis for each fitting parameter in one direction:
Maximum identifiability steps = 50 - The maximal deviation from the initial objective function value is the same as in the example above:
Maximal deviation = 4.2 - Optimization method parameters (the same as in the test_case_2 optimization):
Optimization method name = Particle swarm optimization
Number of itirations = 200
Number of particles = 50 - A possible path to save results of the analysis:
Output path = data/Collaboration/Demo/Data/Temp/Identifiability results (for diagram_2)
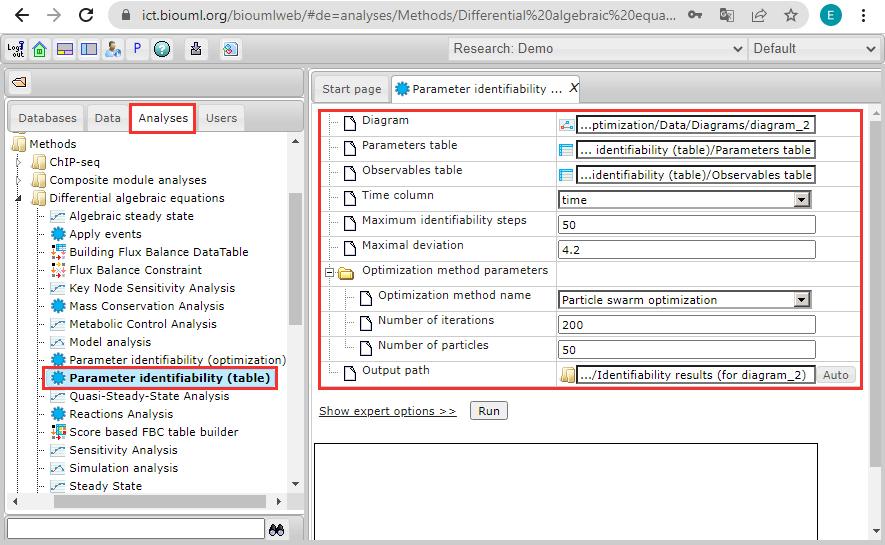
After setting these options, your workspace will look like this:
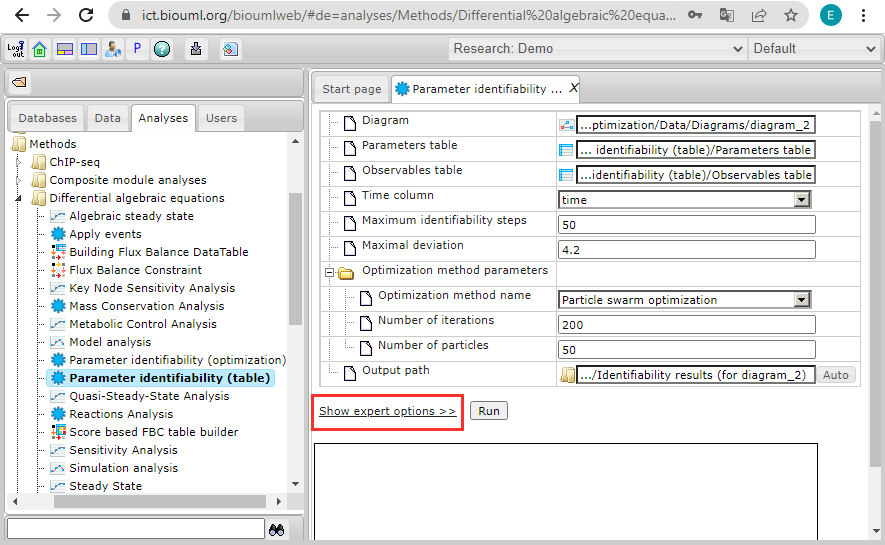
Then click on the phrase "Show expert options >>":
And additionally set the options for complete coincidence with the optimization problem test_case_2:
- The method to estimate the objective function weights:
Weight method = Edited - Path to a table defining the required weights:
Weights table = data/Examples/DAE models/Data/Tables/Parameter identifiability (table)/Weights table

- Name of the column with weights in the Weights table:
Weight column = Weights
To start the analysis, click on the Run button. The results table is similar to the one obtained from the analysis using the optimization document.
[edit] Interpretation of results
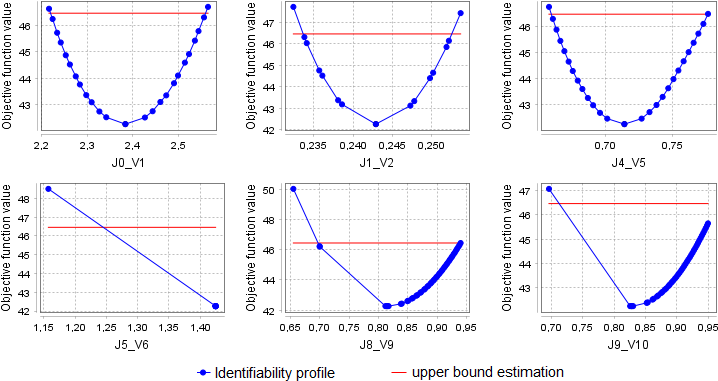
In our example, we received the following identifiability profiles:
Identifiability profiles for parameters J0_V1, J1_V2, J4_V5, J8_V9 cross the threshold of a given deviation from the initial objective function value in both directions. Therefore, the analysis defined these variables as identifiable. The identifiability profiles for parameters J5_V6 and J9_V10 cross the threshold in only one direction, so the analysis determined these parameters as partially non-identifiable. However, it should be noted that the resulting J5_V6 value in the optimization problem is almost equal to the upper bound of the given search space. Thus, the identifiability analysis tried to take very small steps to the right in the parameter value, but this was not enough to significantly change the objective function value:
INFO - (Step 1 of 50) J5_V6 = 1.4249999525529151 (step size = 9.683078561195658E-10), objective function = 42.25511865896442
INFO - (Step 2 of 50) J5_V6 = 1.4249999614126978 (step size = 8.859782795778178E-9), objective function = 42.25511100723304
INFO - (Step 3 of 50) J5_V6 = 1.4249999710878445 (step size = 9.675146601603706E-9), objective function = 42.25509033270342
INFO - (Step 4 of 50) J5_V6 = 1.4249999807495213 (step size = 9.66167670891121E-9), objective function = 42.25508972805858
INFO - (Step 5 of 50) J5_V6 = 1.4249999904319726 (step size = 9.682451299316246E-9), objective function = 42.255089526361225
INFO - (Step 6 of 50) J5_V6 = 1.425 (step size = 9.682869310218694E-9), objective function = 42.25525810172391
INFO - Parameter bound exceeded.
An increase in the J9_V10 value (steps to the right) gave a stable increase in the objective function value. However, the specified threshold was not exceeded because the parameter value reached the upper bound of the search space. Thus, changing the boundaries of the search space in the optimization problem can change the identifiability profile of model parameters.
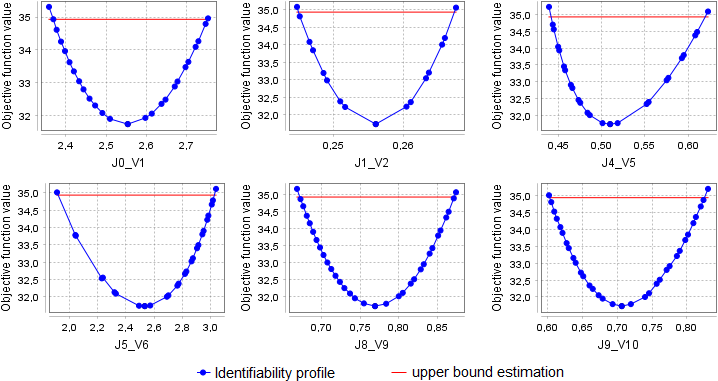
It is also worth noting that the step sizes automatically determined by the algorithm, in particular, depend on how close the value of the fitting parameter is to the search boundary. As can be seen from the identifiability profiles above, for parameters J5_V6, J8_V9, and J9_V10, steps to the right are much smaller than steps to the left. An increase in the upper search bounds for these parameters leads to finding a new solution to the optimization problem (the value of the objective function is 31.7 versus 42.2). In this case, all parameters become identifiable:
[edit] References
- Raue A, Kreutz C, Maiwald T, Bachmann J, Schilling M, Klingmüller U, Timmer J (2009) Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics, 25(15):1923–1929.
- Raue A, Becker V, Klingmüller U, Timmer J (2010) Identifiability and observability analysis for experimental design in nonlinear dynamical models. Chaos, 20(4):045105.
- Kholodenko BN (2000) Negative feedback and ultrasensitivity can bring about oscillations in the mitogenactivated protein kinase cascades. European Journal of Biochemistry, 267(6):1583–1588.