Difference between revisions of "Gene set enrichment analysis - select a classification (Gene table) (workflow)"
(Automatic synchronization with BioUML) |
(Automatic synchronization with BioUML) |
||
| Line 6: | Line 6: | ||
[[File:Gene-set-enrichment-analysis-select-a-classification-Gene-table-workflow-overview.png|400px]] | [[File:Gene-set-enrichment-analysis-select-a-classification-Gene-table-workflow-overview.png|400px]] | ||
== Description == | == Description == | ||
| − | This workflow | + | This workflow is designed to perform Gene Set Enrichment Analysis, GSEA, as it is described at [http://www.broadinstitute.org/gsea/index.jsp http://www.broadinstitute.org/gsea/index.jsp]. As input, any gene or protein table that includes fold change (or any other numerical value that can be used as a weighted column) can be taken. |
| − | + | At the first step, the input table is converted into a table with Ensembl Gene IDs. | |
| − | + | This table with Ensembl Gene IDs is subjected to GSEA. Enrichment analysis is done in parallel by the following ontologies: GO biological processes, GO cellular components, GO molecular functions and by the Reactome pathways. | |
| + | |||
| + | For each ontological term several parameters are calculated, including nominal p-value, ES, NES, as well as hit names, the link to the corresponding ontological term, and the link to open a visualization plot. | ||
== Parameters == | == Parameters == | ||
Revision as of 16:19, 11 December 2014
- Workflow title
- Gene set enrichment analysis - select a classification (Gene table)
- Provider
- geneXplain GmbH
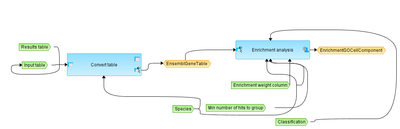
Workflow overview
Description
This workflow is designed to perform Gene Set Enrichment Analysis, GSEA, as it is described at http://www.broadinstitute.org/gsea/index.jsp. As input, any gene or protein table that includes fold change (or any other numerical value that can be used as a weighted column) can be taken.
At the first step, the input table is converted into a table with Ensembl Gene IDs.
This table with Ensembl Gene IDs is subjected to GSEA. Enrichment analysis is done in parallel by the following ontologies: GO biological processes, GO cellular components, GO molecular functions and by the Reactome pathways.
For each ontological term several parameters are calculated, including nominal p-value, ES, NES, as well as hit names, the link to the corresponding ontological term, and the link to open a visualization plot.
Parameters
- Input table
- Enrichment weight column
- Enter the name of the column from the Input table with numerical values
- Species
- Classification
- Min number of hits to group
- Results table