Difference between revisions of "Gene set enrichment analysis - select a classification (Gene table) (workflow)"
m (Protected "Gene set enrichment analysis - select a classification (Gene table) (workflow)": Autogenerated page ([edit=sysop] (indefinite))) |
(Automatic synchronization with BioUML) |
||
| Line 6: | Line 6: | ||
[[File:Gene-set-enrichment-analysis-select-a-classification-Gene-table-workflow-overview.png|400px]] | [[File:Gene-set-enrichment-analysis-select-a-classification-Gene-table-workflow-overview.png|400px]] | ||
== Description == | == Description == | ||
| − | This workflow | + | This workflow performs ''Gene Set Enrichment Analysis'' for the input table of genes or proteins. It is designed to enable a comfortable selection of several input parameters in the input form, especially selection of an ontology. Any ontology available with a given subscription can be chosen from the drop-down menu in the field ''Classification''. |
| − | + | The Input table in the first step is converted to Ensembl Gene IDs, and in the next step an Ensembl gene table is submitted to ''Enrichment analysis'' using the ontology selected. | |
| − | + | The output contains the results of the enrichment analysis. For each ontological term several parameters are calculated, including nominal p-value, ES, NES, Rank at max, FDR, as well as hit names, the link to the corresponding ontological term, and the link to open a visualization plot. | |
| − | + | ||
| − | For each ontological term several parameters are calculated, including nominal p-value, ES, NES, as well as hit names, the link to the corresponding ontological term, and the link to open a visualization plot. | + | |
== Parameters == | == Parameters == | ||
Revision as of 11:49, 30 July 2013
- Workflow title
- Gene set enrichment analysis - select a classification (Gene table)
- Provider
- geneXplain GmbH
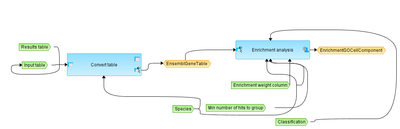
Workflow overview
Description
This workflow performs Gene Set Enrichment Analysis for the input table of genes or proteins. It is designed to enable a comfortable selection of several input parameters in the input form, especially selection of an ontology. Any ontology available with a given subscription can be chosen from the drop-down menu in the field Classification.
The Input table in the first step is converted to Ensembl Gene IDs, and in the next step an Ensembl gene table is submitted to Enrichment analysis using the ontology selected.
The output contains the results of the enrichment analysis. For each ontological term several parameters are calculated, including nominal p-value, ES, NES, Rank at max, FDR, as well as hit names, the link to the corresponding ontological term, and the link to open a visualization plot.
Parameters
- Input table
- Enrichment weight column
- Enter the name of the column from the Input table with numerical values
- Species
- Classification
- Min number of hits to group
- Results table