Difference between revisions of "ChIP-Seq - Identify TF binding sites on peaks (TRANSFAC(R)) (workflow)"
From BioUML platform
(Automatic synchronization with BioUML) |
(Automatic synchronization with BioUML) |
||
| Line 6: | Line 6: | ||
[[File:ChIP-Seq-Identify-TF-binding-sites-on-peaks-TRANSFAC-R-workflow-overview.png|400px]] | [[File:ChIP-Seq-Identify-TF-binding-sites-on-peaks-TRANSFAC-R-workflow-overview.png|400px]] | ||
== Description == | == Description == | ||
| − | This workflow is designed to search for | + | This workflow is designed to search for binding sites for transcription factors on DNA sequences identified by ChiP-Seq approach. Potential binding sites are identified using TRANSFAC(R) database and the results are further refined for significance. |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
== Parameters == | == Parameters == | ||
Latest revision as of 16:19, 11 December 2014
- Workflow title
- ChIP-Seq - Identify TF binding sites on peaks (TRANSFAC(R))
- Provider
- geneXplain GmbH
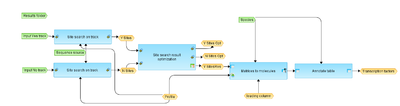
[edit] Workflow overview
[edit] Description
This workflow is designed to search for binding sites for transcription factors on DNA sequences identified by ChiP-Seq approach. Potential binding sites are identified using TRANSFAC(R) database and the results are further refined for significance.
[edit] Parameters
- Input Yes track
- Sequence source
- Species
- Input No track
- Results folder