GTRD
GTRD (Gene Transcription Regulation Database) is a database of transcription factor binding sites identified from ChIP-seq experiments. GTRD analyze freely avalable ChIP-seq experiments from literature, GEO, SRA and ENCODE databases.
The web interface to GTRD is available here.
Database statistics
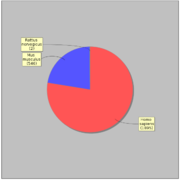
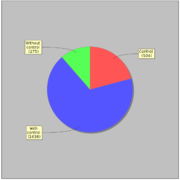
GTRD uses 2417 ChIP-seq experiments for 470 distinct sequence specific transcription factors. Most of ChIP-seq experiments (1638) have corresponding control experiment.General statistics:
| Object type | Total count | Per ChIP-seq experiment |
|---|---|---|
| ChIP-seq reads | 80.808E9 | 34.937E6 |
| Reads aligned | 58.848E9 | 25.675E6 |
| ChIP-seq peaks | 59.515E6 | 32899 |
In average each transcription factor is measured in 4.07 ChIP-seq experiments, but 284 (60%) transcription factors measured only in one experiment.
The ten most studied transcription factors listed bellow:
| Transcription Factor | Number of ChIP-seq experiments |
|---|---|
| CTCF | 195 |
| c-Myc | 45 |
| ERα | 44 |
| NRSF | 37 |
| C/EBPβ | 37 |
| GATA-1 | 33 |
| NF-κB p65 | 30 |
| Max | 30 |
| PU.1 | 29 |
| GR | 24 |
Database structure
The metadata concerning GTRD is stored in MySQL tables.
Each ChIP-seq experiment has a row in 'chip_experiments' table, which assigns id and stores basic information about experiment. 'chip_experiments' table has following structure:
| Column | Description | Example value |
|---|---|---|
| id | Unique experiment identifier | EXP000489 |
| antibody | Antibody used in chromatin immunoprecipitation | sc-345 |
| tfClassId | Id in TFClass[1] database of target transcription factor, NULL for control experiments | 6.2.1.0.1 |
| cell_line | Studied cell line | HeLa S3 |
| specie | Specie latin name | Homo sapiens |
| treatment | Cell treatment or conditions | IFN gamma |
| control_id | Id of control experiment, NULL for control experiments or experiments without control | EXP000490 |
The links to external databases stored in 'external_refs' table:
| Column | Description | Example values |
|---|---|---|
| id | Experiment identifier | EXP000489 |
| external_db | External database name | GEO or PUBMED or ENCODE or SRA |
| external_db_id | Identifier in external database | GSM320736 |
GTRD uses following object identifiers:
| Template | Object type | Example |
|---|---|---|
| EXPXXXXXX | ChIP-seq experiment | EXP000489 |
| READSXXXXXX | Collection of ChIP-seq reads | READS000770 |
| ALIGNSXXXXXX | Collection of read alignments | ALIGNS010001 |
| PEAKSXXXXXX | Collection of ChIP-seq peaks | PEAKS010000 |
The relationship between these objects is provided by 'hub' table:
| Column | Description | Example values |
|---|---|---|
| input | Input object identifier | READS000770 |
| input_type | Type of input object | ReadsGTRDType |
| output | Output object identifier | EXP000489 |
| output_type | Type of output object | ExperimentGTRDType |
ChIP-seq reads, alignments and peaks links to experiments with hub table in the following way:
| input | input_type | output | output_type |
|---|---|---|---|
| READS000770 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000771 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000772 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000773 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000774 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| READS000775 | ReadsGTRDType | EXP000489 | ExperimentGTRDType |
| ALIGNS010001 | AlignmentsGTRDType | EXP000489 | ExperimentGTRDType |
| PEAKS010000 | PeaksGTRDType | EXP000489 | ExperimentGTRDType |
Web interface to database
| This page or section is a stub. Please add screenshots here! |
Web interface to GTRD is available at [2].
It provides capabilities for searching and browsing GTRD.
To search for stat transcription factors enter stat* in the search field and press enter, the list of matching ChIP-seq experiments will appear in the 'Search result' tab. Select one of them to view detailed ChIP-seq experiment information in the 'Info' tab. The 'Info' tab also provides links to the experiment data: reads, alignments and peaks.
For convenience some views are available as ![]() tree-tables.
tree-tables.