Difference between revisions of "Antimony"
(→Composite model) |
Ilya Kiselev (Talk | contribs) |
||
| Line 26: | Line 26: | ||
<table border="1" align="center" cellspacing="0" cellpadding="4"> | <table border="1" align="center" cellspacing="0" cellpadding="4"> | ||
<tr bgcolor="#999999"> | <tr bgcolor="#999999"> | ||
| − | <td> | + | <td> Supported Antimony features </td> |
| − | <td> | + | <td> Not supported in current version</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
<td> | <td> | ||
<ul> | <ul> | ||
| − | <li>Creating and | + | <li>Creating and removing of species, compartments,</li> |
<li>Putting nodes into parental compartments,</li> | <li>Putting nodes into parental compartments,</li> | ||
| − | <li> | + | <li>Initial values changing,</li> |
| − | <li>Creating and | + | <li>Creating and removing of reaction with reactants and products,</li> |
| − | <li>Editing kinetic laws | + | <li>Editing of reaction kinetic laws,</li> |
| − | <li>Creating, | + | <li>Creating, removing and editing of functions,</li> |
| − | <li>Creating, | + | <li>Creating, removing and editing of scalar and rate equations,</li> |
| − | <li> | + | <li>Declarating species and variables as constants;</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 45: | Line 45: | ||
<ul> | <ul> | ||
<li>Declaration <i>"DNA strand definitions"</i> isn't supported,</li> | <li>Declaration <i>"DNA strand definitions"</i> isn't supported,</li> | ||
| − | <li>From symbol types only | + | <li>From symbol types, only "species" and "compartment" are supported,</li> |
| − | <li>Reactions with interactions (inhibition, activation) | + | <li>Reactions with interactions (inhibition, activation);</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 57: | Line 57: | ||
<ul> | <ul> | ||
<li>Algebraic rules</li> | <li>Algebraic rules</li> | ||
| − | <li> | + | <li>Distinguishing between initial value and initial assignment with numerical value,</li> |
| − | <li> | + | <li>Independent management of "constant" and "boundary conditional" attributes,</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 68: | Line 68: | ||
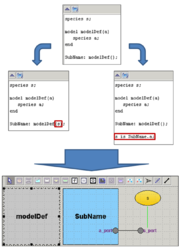
[[File:Ports.png |thumb| Use parameters of subdiagram or synchronization for adding connection.]] | [[File:Ports.png |thumb| Use parameters of subdiagram or synchronization for adding connection.]] | ||
| − | In the beta-version plugin new declarations were added for working with composite models. If antimony text contains few models or declarations outside model then this text | + | In the beta-version plugin new declarations were added for working with composite models. If antimony text contains few models or declarations outside model then this text corresponds to the composite model. Main model will be created and it contains all models like model definitions. |
<table border="1" align="center" cellspacing="0" cellpadding="4"> | <table border="1" align="center" cellspacing="0" cellpadding="4"> | ||
<tr bgcolor="#999999"> | <tr bgcolor="#999999"> | ||
| − | <td> | + | <td> Supported Antimony features </td> |
| − | <td> | + | <td> Not supported in current version</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| Line 81: | Line 81: | ||
<li>External models,</li> | <li>External models,</li> | ||
<li>SubDiagrams,</li> | <li>SubDiagrams,</li> | ||
| − | <li>Replacing | + | <li>Replacing species and parameters from different models;</li> |
</ul> | </ul> | ||
</td> | </td> | ||
<td rowspan="4" valign="top" > | <td rowspan="4" valign="top" > | ||
<ul> | <ul> | ||
| − | <li>Sign < * > for defining main model | + | <li>Sign < * > for defining main model,</li> |
| − | <li>Replacing | + | <li>Replacing of elements with more than one level of depth |
<br><b>Example:</b> s1 is A.B.s2,</li> | <br><b>Example:</b> s1 is A.B.s2,</li> | ||
| − | <li>Replacing elements | + | <li>Replacing of elements other than species and parameters,</li> |
| − | <li> | + | <li>Replacing of elements with different types,</li> |
| − | <li | + | <li>Import models from other collections,</li> |
| − | <li> | + | <li>"Deletion" attributes,</li> |
| + | <li>Operating with submodel elements wich are not defined in model definition as ports explicitly;</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| − | <td bgcolor="#999999">New elements which we added in Antimony language</td> | + | <td bgcolor="#999999">New elements which we have added in Antimony language</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
Revision as of 16:16, 14 November 2013
Contents |
Antimony plugin
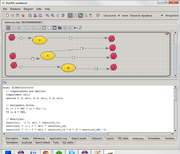
Antimony plugin for BioUML 0.9.6 aims to combine two representations of mathematical model (particularly - SBML model):
- visual representation as BioUML-diagram using extended SBGN notation
- text-based representation using Antimony language[1].
Plugin allows user to edit model both visually and as text.
If visual repesentation is changed - antimony text is updated on the fly.
Text changes are applied to the diagram on demand (after pressing "Apply Antimony" button).
Besides model representations syncrhonisation, we also try to presereve user diagram layout as well as antimony text format (spaces, comments, etc.)
Please note: antimony changes will be applied to the diagram only after "Apply Antimony" button is pressed.
If you change diagram before you press this button, all changes in text will be lost!
Features overview
Currently plugin is in beta version, some features of antimony aren't supported. Also we want to achieve full synchronization between antimony and sbml-model in BioUML. And for this purpose we need some additions for the antimony specification.
| Supported Antimony features | Not supported in current version |
|
|
| Necessary additions in the antimony specification | |
|
Composite model
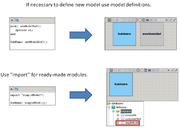
In the beta-version plugin new declarations were added for working with composite models. If antimony text contains few models or declarations outside model then this text corresponds to the composite model. Main model will be created and it contains all models like model definitions.
| Supported Antimony features | Not supported in current version |
|
|
| New elements which we have added in Antimony language | |
|
References
- ↑ Smith, L.P., Bergmann, F.T., Chandran, D. Sauro, M.H. Antimony: a modular model definition language. Bioinformatics, 2009, 25(18): 2452-2454. doi:10.1093/bioinformatics/btp401