Difference between revisions of "ChIP-Seq - Identify and classify target genes (TRANSPATH(R)) (workflow)"
From BioUML platform
m (Protected "ChIP-Seq - Identify and classify target genes (TRANSPATH(R)) (workflow)": Autogenerated page ([edit=sysop] (indefinite))) |
(Automatic synchronization with BioUML) |
||
| Line 11: | Line 11: | ||
;Input track | ;Input track | ||
:Enter your Chip-seq peak track | :Enter your Chip-seq peak track | ||
| + | ;Annotation source | ||
;Species | ;Species | ||
;Results folder | ;Results folder | ||
Latest revision as of 16:35, 12 March 2019
- Workflow title
- ChIP-Seq - Identify and classify target genes (TRANSPATH(R))
- Provider
- geneXplain GmbH
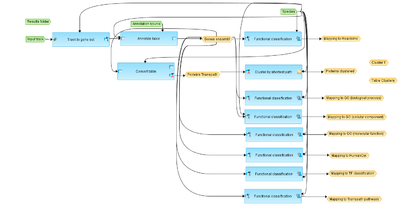
[edit] Workflow overview
[edit] Description
This workflow is designed to identify and classify target genes using positional information of peaks found by ChiP-Seq approach. Classification includes enrichment analysis using Reactome, GeneWays, Proteome GO, HumanCyc , TF classification and Transpath databases.
[edit] Parameters
- Input track
- Enter your Chip-seq peak track
- Annotation source
- Species
- Results folder