Difference between revisions of "Matrix (element type)"
From BioUML platform
Tagir Valeev (Talk | contribs) m (+Description headline) |
Tagir Valeev (Talk | contribs) |
||
| (One intermediate revision by one user not shown) | |||
| Line 4: | Line 4: | ||
Matrix element type represents a [[wikipedia:Position-specific scoring matrix|position weight matrix]] (PWM) which describes some motif. Number of matrix rows depends on the alphabet used (usually 4 rows for mononucleotide alphabet and 16 rows for dinucleotide alphabet). | Matrix element type represents a [[wikipedia:Position-specific scoring matrix|position weight matrix]] (PWM) which describes some motif. Number of matrix rows depends on the alphabet used (usually 4 rows for mononucleotide alphabet and 16 rows for dinucleotide alphabet). | ||
| − | Matrices can only reside inside {{Type link|matrix library}}. Some analyses like [[ChIPMunk (analysis)|ChIPMunk | + | Matrices can only reside inside {{Type link|matrix library|matrix libraries}}. Some analyses like [[ChIPMunk (analysis)|ChIPMunk]] may produce new matrices. |
Matrices themselves do not have any thresholds to predict whether the binding element (transcription factor) binds with given location or not. For this purpose you should use {{Type link|site model}}s. | Matrices themselves do not have any thresholds to predict whether the binding element (transcription factor) binds with given location or not. For this purpose you should use {{Type link|site model}}s. | ||
Matrices cannot be opened as documents. However when you select the matrix in the tree, you can read the comprehend description in the [[information box]]. | Matrices cannot be opened as documents. However when you select the matrix in the tree, you can read the comprehend description in the [[information box]]. | ||
Latest revision as of 17:04, 17 May 2013
[edit] Description
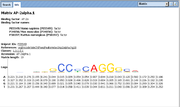
Matrix element type represents a position weight matrix (PWM) which describes some motif. Number of matrix rows depends on the alphabet used (usually 4 rows for mononucleotide alphabet and 16 rows for dinucleotide alphabet).
Matrices can only reside inside ![]() matrix libraries. Some analyses like ChIPMunk may produce new matrices.
matrix libraries. Some analyses like ChIPMunk may produce new matrices.
Matrices themselves do not have any thresholds to predict whether the binding element (transcription factor) binds with given location or not. For this purpose you should use ![]() site models.
site models.
Matrices cannot be opened as documents. However when you select the matrix in the tree, you can read the comprehend description in the information box.