Difference between revisions of "ChIP-Seq - Identify TF binding sites on peaks (TRANSFAC(R)) (workflow)"
m (Cosmetic changes in category formatting) |
(Automatic synchronization with BioUML) |
||
| Line 6: | Line 6: | ||
[[File:ChIP-Seq-Identify-TF-binding-sites-on-peaks-TRANSFAC-R-workflow-overview.png|400px]] | [[File:ChIP-Seq-Identify-TF-binding-sites-on-peaks-TRANSFAC-R-workflow-overview.png|400px]] | ||
== Description == | == Description == | ||
| − | This workflow is designed to search for | + | This workflow is designed to search for TFBSs on DNA sequences identified by the ChiP-Seq approach. Actually, any dataset in BED format can be submitted as input track for this workflow. |
| + | |||
| + | As input, two different tracks should be submitted, one is a dataset under study, Yes track, and another is a background dataset, No track. The default No track corresponds to far upstream regions of the house keeping genes, where no functional composite modules are expected. | ||
| + | |||
| + | In the first step both Yes and No tracks are submitted to the ''Site search on track'' and ''Site search results optimization'' analyses to find TFBSs enriched in the Yes-track as compared with the No-track. The workflow uses the default profile vertebrate_non_redundant_minSUM from the TRANSFAC® library. The output of this step is a list of PWMs the hits of which are enriched in Yes set versus No set. | ||
| + | |||
| + | Next, the list of PWMs is converted into a table of genes with Ensembl IDs using the ''Matrices to molecules'' analysis, and further annotated with additional information, gene descriptions and gene symbols via ''Annotate table'' analysis. | ||
| + | |||
| + | This workflow is available together with a valid TRANSFAC® license | ||
| + | |||
| + | |||
== Parameters == | == Parameters == | ||
Revision as of 11:49, 30 July 2013
- Workflow title
- ChIP-Seq - Identify TF binding sites on peaks (TRANSFAC(R))
- Provider
- geneXplain GmbH
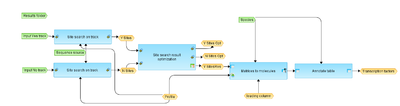
Workflow overview
Description
This workflow is designed to search for TFBSs on DNA sequences identified by the ChiP-Seq approach. Actually, any dataset in BED format can be submitted as input track for this workflow.
As input, two different tracks should be submitted, one is a dataset under study, Yes track, and another is a background dataset, No track. The default No track corresponds to far upstream regions of the house keeping genes, where no functional composite modules are expected.
In the first step both Yes and No tracks are submitted to the Site search on track and Site search results optimization analyses to find TFBSs enriched in the Yes-track as compared with the No-track. The workflow uses the default profile vertebrate_non_redundant_minSUM from the TRANSFAC® library. The output of this step is a list of PWMs the hits of which are enriched in Yes set versus No set.
Next, the list of PWMs is converted into a table of genes with Ensembl IDs using the Matrices to molecules analysis, and further annotated with additional information, gene descriptions and gene symbols via Annotate table analysis.
This workflow is available together with a valid TRANSFAC® license
Parameters
- Input Yes track
- Sequence source
- Species
- Input No track
- Results folder