TRRD
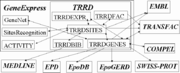
The Transcription Regulatory Regions Database (TRRD) was designed as a convenient tool for studying the complex process of transcription regulation. The model of structure-function organization of transcription regulatory regions of eukaryotic genes that formed the basis of the TRRD database considers a great diversity of elements involved in transcription control, their block structure, and the hierarchy essential for their functioning. In its 4th release (1998) the TRRD database contained five interconnected tables:
- TRRDGENES (general description of genes),
- TRRDEXP (description of expression patterns),
- TRRDSITES (description of sites),
- TRRDFAC (description of transcription factors), and
- TRRDBIB (references to original papers).
In addition, the table TRRDGENES was furnished with the references to EMBL, SwissProt, EPD, EpoGERD, and EpoDB databases as well as to the GeneNet, a section of GeneExpress.
The table TRRDSITES had the references to EMBL and TRANSFAC databases as well as to site recognition programs and the ACTIVITY database, included in the GeneExpress system.
The table TRRDFAC contains the references to TRANSFAC; the table TRRDBIB, to MEDLINE.
The 4th release of TRRD (1998) comprised descriptions of about 500 genes, over 800 regulatory units (promoters, enhancers, and silencers), and about 4047 transcription factor binding sites. Over 1500 scientific publications were processed to obtain these data. The genes described in TRRD belong to different eukaryotic species. Human (41%) and mouse (25%) genes constitute the major part; rat (15%) and chick genes were well represented too.
The TRRD database was a part of the global system GeneExpress available at the Molecular Biological Server of the Institute of Cytology and Genetics, Siberian Branch of the Russian Academy of Sciences.
TRRD-Viewer
The applet TRRD-Viewer allowed for visualization of the data on the location of transcription factor binding sites in a map form and displayed their textual description.
When working with this applet, the user would select a gene identifier from the list, and the textual description of the gene (from TRRDGENES), its sites (from TRRDSITES), and the relevant references (from TRRDBIB) would appear in the text window, while the transcription factor binding sites and composite elements would be presented graphically. If the user clicked on the site image, its description from the table TRRDSITES was displayed in the text window. Clicking the field title provided the display of the comments on the information described in the field. The applet provided options allowing different site representations.
The work was supported by the Russian Foundation for Basic Research (96-04-50006, 97-04-49740, 98-04-49479), Integration Program of the Siberian Branch of the Russian Academy of Sciences (IGSBRAS-97*13), and Young Scientists Competition of SB RAS.