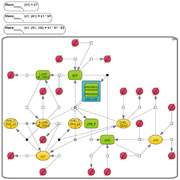
Cross cost grid layout
From BioUML platform
- Title
- Cross cost grid layout
- Class
CompartmentCrossCostGridLayouter
| This page or section is a stub. Please add algorithm details and references here! |
This is a grid-based algorithm that considers (a) edge-edge crossings, (b) node-edge crossings, (c) node-node crossings, (d) distances between nodes in its cost function . This algorithm uses a weight matrix representing the difference between two sequentially obtained layouts for computing the costs, takes a greedy algorithm for searching locally optimal solutions and simulated annealing for global optimization.
The following parameters of the algorithm must be set:
- GridX - positive integer parameter represents horizontal grid step in pixels
- GridY - positive integer parameter represents vertical grid step in pixels
- Number of iteration - positive integer parameter represents number of iterations during same temperature in simulated annealing
- Cooling coefficient - real number between 0 and 1 represents cooling coefficient of simulated annealing (small value decrease computation time but leads to quality loss)
- Perturbation threshold - real value parameter between 0 and 1 represents probability of layout perturbation during simulated annealing
- Max distance - positive integer parameter represents maximum repulsive distance expressed in number of grid steps (if distance between two nodes exceeds Max distance then there is no repulsion between them).