Antimony
Contents |
Antimony plugin
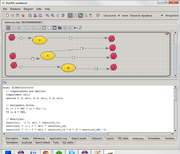
Antimony plugin for BioUML 2023.3 aims to combine two representations of mathematical model (particularly - SBML model):
- visual representation as BioUML
 diagram using extended SBGN notation
diagram using extended SBGN notation
- text-based representation using Antimony language[1].
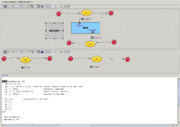
Plugin allows user to edit model both visually and as text.
If visual repesentation is changed - antimony text is updated on the fly.
Text changes are applied to the ![]() diagram on demand (after pressing "Apply Antimony" button).
diagram on demand (after pressing "Apply Antimony" button).
Besides model representations syncrhonisation, we also try to presereve user diagram layout as well as antimony text format (spaces, comments, etc.)
Please note: antimony changes will be applied to the ![]() diagram only after "Apply Antimony" button is pressed.
If you change
diagram only after "Apply Antimony" button is pressed.
If you change ![]() diagram before you press this button, all changes in text will be lost!
diagram before you press this button, all changes in text will be lost!
Features overview
Currently plugin is in beta version, some features of antimony aren't supported. Also we want to achieve full synchronization between antimony and sbml-model in BioUML. And for this purpose we need some additions for the antimony specification.
| Supported Antimony features | Not supported in current version |
|
|
| Necessary additions in the antimony specification | |
|
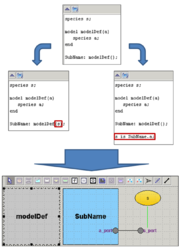
Composite model
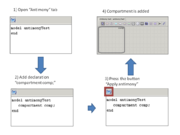
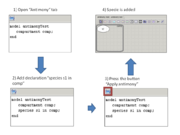
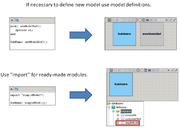
In the beta-version plugin new declarations were added for working with composite models. If antimony text contains few models or declarations outside model then this text corresponds to the composite model. Main model will be created and it contains all models like model definitions.
| Supported Antimony features | Not supported in current version |
|
|
| New elements which we have added in Antimony language | |
|
Other features
Attributes. Antimony language has been expended for working with element's attributes. String attribute can be specified by declaration: @elementName.attributeName = "value". Some attributes are not displayed in the antimony text, but one is able to choose visible attributes in "Additional properties" (tab Options). List of attributes is split by commas.
References
- ↑ Smith, L.P., Bergmann, F.T., Chandran, D. Sauro, M.H. Antimony: a modular model definition language. Bioinformatics, 2009, 25(18): 2452-2454. doi:10.1093/bioinformatics/btp401