Difference between revisions of "Find master regulators for multiple gene sets (TRANSPATH(R)) (workflow)"
(Automatic synchronization with BioUML) |
m (Cosmetic changes in category formatting) |
||
| (4 intermediate revisions by one user not shown) | |||
| Line 2: | Line 2: | ||
:Find master regulators for multiple gene sets (TRANSPATH(R)) | :Find master regulators for multiple gene sets (TRANSPATH(R)) | ||
;Provider | ;Provider | ||
| − | :[[ | + | :[[geneXplain GmbH]] |
== Workflow overview == | == Workflow overview == | ||
[[File:Find-master-regulators-for-multiple-gene-sets-TRANSPATH-R-workflow-overview.png|400px]] | [[File:Find-master-regulators-for-multiple-gene-sets-TRANSPATH-R-workflow-overview.png|400px]] | ||
| Line 31: | Line 31: | ||
[[Category:Workflows]] | [[Category:Workflows]] | ||
| + | [[Category:GeneXplain workflows]] | ||
[[Category:Autogenerated pages]] | [[Category:Autogenerated pages]] | ||
Latest revision as of 13:28, 16 May 2013
- Workflow title
- Find master regulators for multiple gene sets (TRANSPATH(R))
- Provider
- geneXplain GmbH
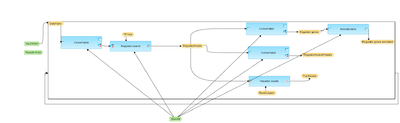
[edit] Workflow overview
[edit] Description
This workflow is designed to find master regulatory molecules upstream of an input list of genes. The Input is a folder containing several gene or protein tables, and these tables are taken automatically by the workflow, one input table after another, in a cycle.
At the first step, one of the input tables from the input folder is converted into TRANSPATH® peptides.
The table with TRANSPATH® peptides is subjected to master regulator search in the TRANSPATH® network. For each potential master regulator, FDR, Score, and Z-score are calculated.
The master regulatory molecules are filtered by Z_Score>1 and Score>0.2 to select statistically significant master regulators.
At the next step, the filtered regulatory molecules are converted to both Ensembl Gene IDs and a list of UniProt protein IDs.
The table with Ensembl Gene Ids is annotated with additional information, gene description and gene symbols.
Finally, the table with master regulatory molecules is sorted by Score and networks for the three top master regulators are visualized as diagrams in the hierarchical layout for all input tables.
The same steps are repeated for the next input table, and several cycles are performed automatically corresponding to the number of tables in the input folder.
This workflow is available together with a valid TRANSPATH® license.
[edit] Parameters
- Input folder
- Input the folder containing differentially expressed genes
- Species
- Results folder