Difference between revisions of "Quantification of RNA-seq with Cufflinks (with de-novo assembly) for FASTQ files (workflow)"
(Automatic synchronization with BioUML) |
(Automatic synchronization with BioUML) |
||
| (One intermediate revision by one user not shown) | |||
| Line 5: | Line 5: | ||
== Workflow overview == | == Workflow overview == | ||
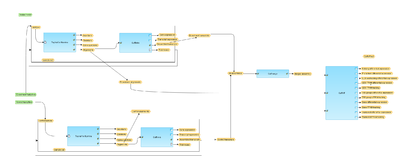
[[File:Quantification-of-RNA-seq-with-Cufflinks-with-de-novo-assembly-for-FASTQ-files-workflow-overview.png|400px]] | [[File:Quantification-of-RNA-seq-with-Cufflinks-with-de-novo-assembly-for-FASTQ-files-workflow-overview.png|400px]] | ||
| + | == Description == | ||
| + | This workflow offers the ability to discover new genes and transcripts (splice variants) and measure transcript expression in a single assay from RNA-seq data. | ||
| + | |||
| + | '''This workflow is described in “Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks”, Nat. Protoc. 7:562-578, 2012.''' | ||
| + | |||
| + | The first step of the workflow is a read alignment with '''TopHat''' (http://tophat.cbcb.umd.edu/). TopHat aligns reads to the genome and discovers transcript splice sites. Output files from TopHat are tables and tracks with insertions, deletions, splice junctions and the alignments. | ||
| + | |||
| + | These output files are subjected to Cufflinks. Cufflinks uses the alignments to map reads against the genome and to assemble the reads into transcripts. Output tracks of Cufflinks is the Assembled transcripts track, output tables of Cufflinks are Gene expression and Transcript expression tables. | ||
| + | |||
| + | In the current workflow the transcripts are assembled “de-novo”, since here it is a “de-novo” reconstruction of exon-intron structure, no known gene or transcript names are given. All transcripts are defined by the tracking_id, like Cuff.1.1 and so on. This allows us to find new transcripts that were not yet discovered and annotated in the reference genome. | ||
| + | |||
| + | At the next step the output of Cufflinks is subjected to Cuffmerge, is essentially a ‘meta-assembler’ — it treats the assembled transfrags the way Cufflinks treats reads, merging them together parsimoniously. Output is a Merged assembly track. | ||
| + | |||
| + | Differential expression is performed by Cuffdiff, part of the Cufflinks package (http://cufflinks.cbcb.umd.edu/), which calculates expression in two or more samples and tests the statistical significance of each observed change in expression between them. | ||
| + | |||
| + | Output is a folder with many differentially expressed genes. | ||
| + | |||
| + | |||
| + | |||
== Parameters == | == Parameters == | ||
;Experiment fastq files | ;Experiment fastq files | ||
;Control fastq files | ;Control fastq files | ||
| − | |||
| − | |||
;Output folder | ;Output folder | ||
Latest revision as of 16:34, 12 March 2019
- Workflow title
- Quantification of RNA-seq with Cufflinks (with de-novo assembly) for FASTQ files
- Provider
- geneXplain GmbH
[edit] Workflow overview
[edit] Description
This workflow offers the ability to discover new genes and transcripts (splice variants) and measure transcript expression in a single assay from RNA-seq data.
This workflow is described in “Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks”, Nat. Protoc. 7:562-578, 2012.
The first step of the workflow is a read alignment with TopHat (http://tophat.cbcb.umd.edu/). TopHat aligns reads to the genome and discovers transcript splice sites. Output files from TopHat are tables and tracks with insertions, deletions, splice junctions and the alignments.
These output files are subjected to Cufflinks. Cufflinks uses the alignments to map reads against the genome and to assemble the reads into transcripts. Output tracks of Cufflinks is the Assembled transcripts track, output tables of Cufflinks are Gene expression and Transcript expression tables.
In the current workflow the transcripts are assembled “de-novo”, since here it is a “de-novo” reconstruction of exon-intron structure, no known gene or transcript names are given. All transcripts are defined by the tracking_id, like Cuff.1.1 and so on. This allows us to find new transcripts that were not yet discovered and annotated in the reference genome.
At the next step the output of Cufflinks is subjected to Cuffmerge, is essentially a ‘meta-assembler’ — it treats the assembled transfrags the way Cufflinks treats reads, merging them together parsimoniously. Output is a Merged assembly track.
Differential expression is performed by Cuffdiff, part of the Cufflinks package (http://cufflinks.cbcb.umd.edu/), which calculates expression in two or more samples and tests the statistical significance of each observed change in expression between them.
Output is a folder with many differentially expressed genes.
[edit] Parameters
- Experiment fastq files
- Control fastq files
- Output folder