Difference between revisions of "Compute differentially expressed genes using Hypergeometric test (Illumina probes) (workflow)"
m (Cosmetic changes in category formatting) |
(Automatic synchronization with BioUML) |
||
| Line 6: | Line 6: | ||
[[File:Compute-differentially-expressed-genes-using-Hypergeometric-test-Illumina-probes-workflow-overview.png|400px]] | [[File:Compute-differentially-expressed-genes-using-Hypergeometric-test-Illumina-probes-workflow-overview.png|400px]] | ||
== Description == | == Description == | ||
| − | This workflow is designed to identify | + | This workflow is designed to identify differentially expressed genes from an experiment data set compared to a control data set. It can be used even for the case with one data point for each the experiment and the control. Normalized data with Illumina probeset IDs can be submitted as input. Such normalized files are the output of the workflow [http://test.genexplain.com/bioumlweb/#de=analyses/Methods/Data normalization/Illumina normalization Illumina normalization]. |
| − | + | In the first step, the up- and down-regulated probes are identified and log fold change values are calculated for all probes using the ''Fold Change calculation ''analysis. This workflow applies ''Hypergeometric analysis'' for the p-value calculation. | |
| − | + | In addition the results are filtered by different conditions in parallel applying the ''Filter table'' method, to identify up-regulated and down-regulated Agilent probeset IDs. The filtering criteria are set as follows: | |
| − | + | '''For up-regulated probes: LogFoldChange>0.5 and -log_P_value_>3. | |
| − | + | For down- regulated probes: LogFoldChange<-0.5 and -log_P_value_<-3. | |
| − | + | ''' | |
| − | The | + | The resulting tables of up-regulated, down-regulated, and non-changed Illumina probeset IDs are converted into a gene table with the ''Convert table'' method and annotated with additional information (gene descriptions, gene symbols, and species) via ''Annotate table'' method. |
| − | + | A result folder is generated and automatically named corresponding to the experiment data set name. This resulting folder contains all generated tables. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
== Parameters == | == Parameters == | ||
Latest revision as of 16:34, 12 March 2019
- Workflow title
- Compute differentially expressed genes using Hypergeometric test (Illumina probes)
- Provider
- geneXplain GmbH
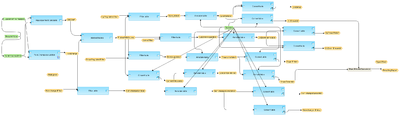
[edit] Workflow overview
[edit] Description
This workflow is designed to identify differentially expressed genes from an experiment data set compared to a control data set. It can be used even for the case with one data point for each the experiment and the control. Normalized data with Illumina probeset IDs can be submitted as input. Such normalized files are the output of the workflow normalization/Illumina normalization Illumina normalization.
In the first step, the up- and down-regulated probes are identified and log fold change values are calculated for all probes using the Fold Change calculation analysis. This workflow applies Hypergeometric analysis for the p-value calculation.
In addition the results are filtered by different conditions in parallel applying the Filter table method, to identify up-regulated and down-regulated Agilent probeset IDs. The filtering criteria are set as follows:
For up-regulated probes: LogFoldChange>0.5 and -log_P_value_>3.
For down- regulated probes: LogFoldChange<-0.5 and -log_P_value_<-3.
The resulting tables of up-regulated, down-regulated, and non-changed Illumina probeset IDs are converted into a gene table with the Convert table method and annotated with additional information (gene descriptions, gene symbols, and species) via Annotate table method.
A result folder is generated and automatically named corresponding to the experiment data set name. This resulting folder contains all generated tables.
[edit] Parameters
- Experiment normalized
- Control normalized
- Species
- Results folder